Yhe authors of Explore Evolution reveal a major gap in their understanding of phylogeny and of much of modern biology when they state:
if Darwin's single Tree of Life is accurate, then we should expect that different types of biological evidence would all point to that same tree. A "family history" of organisms based on their anatomy should match the "family history" based on their molecules (such as DNA and proteins).Explore Evolution, p. 57
Phylogenetic trees based on a specific gene (gene trees) and those based on several genetic and anatomical traits map the relationships of different entities, genes and organisms. Inconsistencies in phylogenetic tree reconstructions are a fascinating issue and research addressing these inconsistencies has led to a better understanding of complex evolutionary processes.
Phylogenetic trees reconstructed from different genes in the same organism can differ. The possible causes of such differences are understood, ranging from methodological issues (such as different parameters being applied to the algorithms used to weigh sequence similarities) to bona fide biological phenomena. The latter are more interesting and significant, and are generally due to the effects of recognized evolutionary processes on the history of individual genes: convergence, the same sequence change appearing independently in different lineages either because of similar selective pressures, or by chance; changes in evolutionary rates , certain organisms evolve faster than others; horizontal gene transfer, sequences being transfered from one species to another by mechanisms other than vertical, linear descent; and timing, two lineages radiate from a third in relatively close succession, before enough differences mayhave accumulated between them to be able to discern the order of emergence. Thus, even in the best scenarios, absolutely congruent phylogenies from the analysis of individual genes are not expected. The authors of Explore Evolution make it seem as if biologists are surprised and stumped by these inconsistencies:
Evolutionary biologist Michael Lynch has noted that creating a clear picture of evolutionary relationships is "an elusive problem." He also notes that "analyses based on different genes - and even different analyses based on the samegenes [yield] a diversity of phylogenetic trees."Explore Evolution, p. 57
But in the very next paragraph of the same paper, Lynch makes the main underlying issues clear:
Given the substantial evolutionary time separating the animal phyla, it is not surprising that single-gene analyses yield such discordant results. Under such circumstances, the statistical noise associated with the substitution process leads to a high probability that phylogenetic analyses based on different molecules will yield different topologies (Philippe et al. 1994; Ruvolo 1997), so that inferences based on single genes can potentially be very misleading (leaving aside for now the additional problem of orthology).[emphasis added]Lynch M. "The Age and Relationships of the Major Animal Phyla." Evolution. 1999; 53:319-325.
In Lynch's paper the phrase "elusive problem" and the issue of multiple phylogenetic trees applied specifically to the "phylogenetic relationships of the major animal phyla", i.e. very distant evolutionary events; the Explore Evolution authors craftily presented it as if Lynch was referring to all "evolutionary relationships".
Another example of the issues encountered in phylogenetic reconstruction, and their misrepresentation in Explore Evolution, comes from the following paragraph:
A "family tree" based on anatomy may show one pattern of relationships, while a tree based on DNA or RNA may show quite another. For example, one analysis of the mitochondrial cytochrome b gene produced a "family tree" in which cats and whales wound up in the order Primates. Yet, an anatomical analysis says that cats belong to the order Carnivora, while whales belong to Cetacea and neither of them are Primates.Explore Evolution, p. 57
The authors are talking about a review paper by Michael Lee (Lee MSY, 1999 Trends Ecol Evol 14:177-178), in which he refers to data obtained on one of the proteins involved in the respiratory chain in mitochondria, cytochrome b. The figure below shows the tree as presented in Lee's review paper: 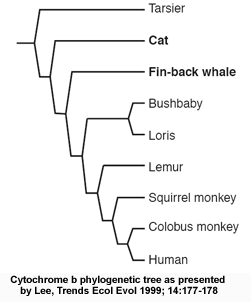 Cytochrome b phylogenetic tree: from Lee, (1999) Trends Ecol Evol 14:177-178
Cytochrome b phylogenetic tree: from Lee, (1999) Trends Ecol Evol 14:177-178
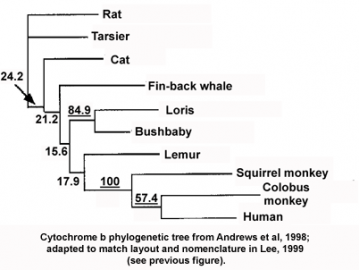
The phylogenetic inconsistency here is the misplacement of a single branch, that of tarsiers (a primitive group of primates), as if they had separated from other primates before cats and fin-back whales. Actually, the data in the original publication (see figure below, Andrews et al. 1998 "Accelerated Evolution of Cytochrome b in Simian Primates: Adaptive Evolution in Concert with Other Mitochondrial Proteins?" J Mol Evol. 47:249 257) gives a slightly different picture, namely that the analysis of cytochrome b sequence is statistically incapable of resolving the phylogenetic relationship of most of the species in the tree (the numbers in the figure represent a measure of the statistical confidence in each branch of the tree, and numbers below 30 generally indicate lower confidence; the statistically robust values are underlined). In other words, cytochrome b is simply not a good protein to choose for constructing the evolutionary tree of these species. But why is that?
 Cytochrome b phylogenetic tree: from Andrews et al., 1998; adapted to match layout and nomenclature in Lee, 1999 (see prevoius figure)
Cytochrome b phylogenetic tree: from Andrews et al., 1998; adapted to match layout and nomenclature in Lee, 1999 (see prevoius figure) Both the Andrews and Lee papers suggested, based on other data, that the phylogenetic incongruence in this tree was caused by cytochrome b and other respiratory chain proteins having evolved much faster in some primate lineages compared to other mammals, possibly following unique selective pressures. As mentioned above, both accelerated and adaptive evolution can cause errors in phylogenetic tree reconstruction, masking or enhancing the similarities of related genes, depending on the circumstances. And indeed, in more recent years the accelerated adaptive evolution of respiratory chain proteins in monkeys and apes (but not tarsiers and lemurs) has been extensively confirmed (see for instance Grossman LI, et al. 2004 "Accelerated evolution of the electron transport chain in anthropoid primates." Trends Genet. 20:578-585). Thus, the inconsistency in the cytochrome b tree, rather than highlighting hopeless phylogenetic confusion as alleged in Explore Evolution, is the result of real biological and evolutionary processes. The existence of this extensive literature offers opportunities for an inquiry-based lesson on molecular evolution and evolutionary processes. Instead of offering that lesson, the supposedly inquiry-based Explore Evolution throws up its hands in confusion at any sign of difficulty.
Although molecular phylogenetic tree inconsistencies are hardly a fundamental theoretical concern for evolutionary biology, if persistent they could still cause practical problems in assessing certain evolutionary relationships. However, a number of new approaches have recently emerged that address these difficulties. These methods include the combination of large sets of sequence information from genomic databases, as well as the use of genetic features, such as large-scale structural changes or the mapping of mobile genetic elements, that are less prone to convergence and selection-related artifacts. For a thorough discussion of the potential of these approaches, see Lokas A and Carroll SB, (2006) "Bushes in the Tree of Life" PLoS Biol 4:e352.
Finally, the authors of Explore Evolution conclude:
Critics point out that the real problem may be that Universal Common Descent is wrong. In other words, maybe the reason the family trees don't agree is that the organisms in question never did share a common ancestor. Even some evolutionary biologists agree. Carl Woese of the University of Illinois, for instance, now thinks that biology must abandon what he calls Darwin's "Doctrine of Common Descent".Explore Evolution, p. 58
Woese argues that the earliest history of life may show multiple early lineages which swapped genes extensively, making reconstruction of the early tree of life difficult. This is very different from the strictly non-overlapping trees Explore Evolution suggests as an alternative to universal common ancestry. Woese argues that these multiple lineages converged into a single population from which modern life, and would absolutely reject the claim that molecular data cannot discern the pattern of common ancestry linking all primates, or the relationship between primates, carnivores, and whales, or indeed the common ancestry of all multicellular organisms.
